For many of us, food is something we buy at a supermarket or order at a café. We usually give little thought to the complex systems required to produce and deliver it—until they stop working. It’s not common to think of Australia as a place at risk of food insecurity. It has vast tracts of fertile land and the capacity to feed its population many times over. Around 70% is exported.
But the searing southeast heat and widespread northern flooding this summer demonstrate the very real risks to food production. Temperature extremes, heat waves, droughts, floods, and shifting seasonal patterns are worsening as the climate changes.
People can seek refuge indoors. But the plants and animals we rely on for food have no such protection. In response, some orchardists and farmers are taking up an approach known as protected cropping, where crops are shielded from threats. As South Australian persimmon and avocado grower Craig Burne told the ABC:
“Without misting and netting in place, I don’t think we’d successfully grow either of these crops in this climate any more.”
As climate change intensifies, protected cropping could better safeguard some crops. Overseas, nations such as the Netherlands have taken up protected cropping to drastically boost fruit and vegetable exports. But it’s early days in Australia. To grow, the sector will have to overcome barriers to growth.
What defines protected cropping?
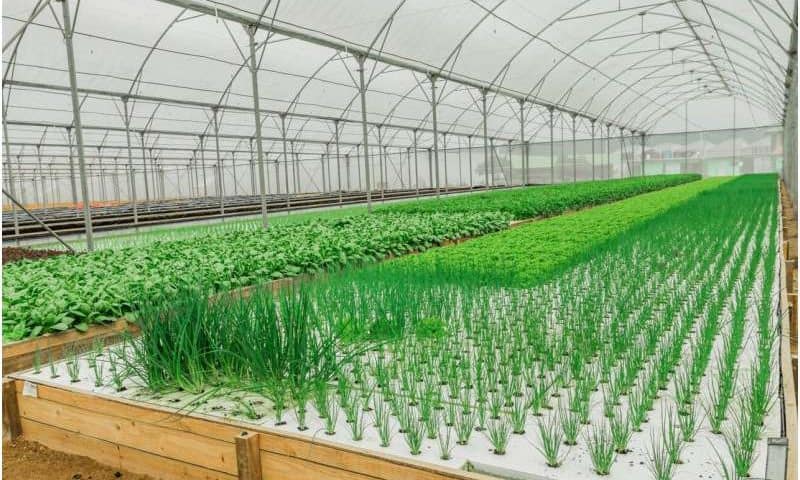
Protection is broadly defined. It can range from low-tech solutions such as shade houses and netting to medium-technology polytunnels (hoop-shaped plastic covers) through to highly sophisticated automated glasshouses.
Countries facing land constraints such as the Netherlands have been the most enthusiastic in taking up this approach. Guided by the principle of “twice the food using half the resources,” Netherlands farmers have turned to high-tech glasshouses.
The result has been remarkable: a country with extremely limited agricultural land has become a top exporter of fruit and vegetables.
Emerging in Australia
In Australia, protected cropping is gaining popularity off a small base. In 2023, around 14,000 hectares of fruit and vegetable crops were growing under some form of protection. That’s around 17% of the total area.
Most of this area relies on low-tech systems, however. Just over two-thirds (68%) of all protected cropping areas rely on low-tech shade houses or netting, mainly in southern Queensland and northern New South Wales.
Medium-tech systems such as polytunnels and polyhouses account for about 30% of the total. These systems are found mainly in Tasmania, northern Queensland, and Western Australia.
High-tech glasshouses account for only 2% of the total. These are primarily found near bigger cities such as Sydney, Melbourne, and Adelaide.
To date, farmers have relied on protected cropping for high-value crops such as tomatoes, capsicums, cucumbers, berries, leafy greens, and more expensive tree crops.
In 2022, Australia’s protected cropping industry was worth an estimated A$100 million to farmers. Demand for workers in the sector is growing at 5% a year, and around 10,000 people worked in the industry as of 2022.
Real benefits—at a cost
For farmers, protected cropping offers clear advantages across low-, medium-, and high-tech approaches.
These methods can create an environment favorable to year-round plant growth, improving the consistency and quality of yields. By controlling factors such as temperature, plant nutrition, humidity, light, and pests, protected cropping reduces production risks and increases crop yield and quality.
For farmers, being able to control their environment in a predictable way is particularly valuable in an uncertain climate. Protecting crops means less (but not zero) risk from extreme weather. Other benefits include more efficient use of land, water, fertilizer and energy.
Crops can also be cultivated closer to markets. This improves food freshness, lowers transport emissions, and strengthens domestic food security.
For exporters, produce grown in protected systems is more likely to meet stringent biosecurity and quality standards of overseas buyers.
Innovation is essential to unlock these benefits at scale. Advances in plant breeding, sensors, automation, data analytics, controlled supply of nutrients, lighting systems, and biological controls for pests and plant diseases can significantly boost farm production, profits, and sustainability.
What’s stopping protected cropping?
Australia’s farmers are highly exposed to extreme weather events and the changing water cycle. Protected cropping would seem to be a logical way to control some of these risks.
To date, protected cropping hasn’t achieved scale in Australia. That’s because the horticulture industry is dominated by small businesses with limited capacity to invest in new systems.
High-tech protected cropping systems offer the best results, but the cost is enough to put off many farmers. Finding and keeping skilled workers is another challenge.
Scaling up won’t just happen
Protected cropping is an excellent solution. But it’s out of reach for many farmers who would benefit.
In nations such as Sweden and the Netherlands, governments have worked to encourage the uptake of protected cropping and boost exports of fruit and vegetables, through world-class research and innovation precincts.
Australia’s federal and state governments could accelerate uptake by setting targets to expand protected cropping areas, encourage adoption through policy levers, investing in joint infrastructure and incentives to cut installation costs.
A good start could be to focus on areas where high-value crops are grown in unprotected environments and work to create regional clusters of expertise, shared infrastructure, and skilled jobs.
Governments can’t do it without buying-in from industry bodies, researchers, and farmers. Translating innovation from laboratory to field is never easy. But it can—and arguably must—be done, as Australia’s farmers face a very uncertain climate.
Protected cropping is not a silver bullet. Polytunnels can’t protect against floods, for instance. But other countries have successfully used these methods to boost yields, safeguard local food production, and create new higher wage jobs. It could do the same here.