AI-powered forecasts sharpen early warning for destructive crop pest
What if farmers could see a pest outbreak coming before the insect ever had a chance to damage their crop? New research from Texas A&M AgriLife Research indicates that artificial intelligence can predict outbreaks much more accurately than traditional methods. The tool could dramatically improve how and when insect pest risks are identified and controlled.
In their study recently published in Ecological Informatics, scientists in the Texas A&M College of Agriculture and Life Sciences Department of Entomology used machine learning models to forecast populations of western flower thrips with notable accuracy, offering producers an early warning when pest pressure is building.
The research was led by Kiran Gadhave, Ph.D., AgriLife Research entomologist and assistant professor in the Department of Entomology at the Texas A&M AgriLife High Plains Research and Extension Center in Canyon. “If we can see pest risk building even a week earlier, that changes everything,” Gadhave said. “Accurately predicting risks sooner shifts management from reacting to damage to staying ahead of it.”
Postdoctoral researcher Arinder Arora, Ph.D., in the Department of Entomology, and Nolan Anderson, Ph.D., AgriLife Research plant pathologist in the Texas A&M Department of Plant Pathology and Microbiology, both at the Texas A&M AgriLife center in Canyon, contributed to the study.
Thousands of traps, millions of data points
Western flower thrips, tiny insects that feed on plants and spread damaging viruses, can trigger significant losses in vegetable and commodity crops once populations begin to surge. By the time producers notice the damage, the outbreak is often underway.
They damage plants during feeding and are considered a “supervector” because even small populations can trigger large yield losses once virus transmission begins.
Traditional forecasting methods for insect pests like thrips in production settings rely on simple parameters, such as temperature, humidity and the number of pests present. But those approaches often fall short in accurately assessing pests’ threat potential.
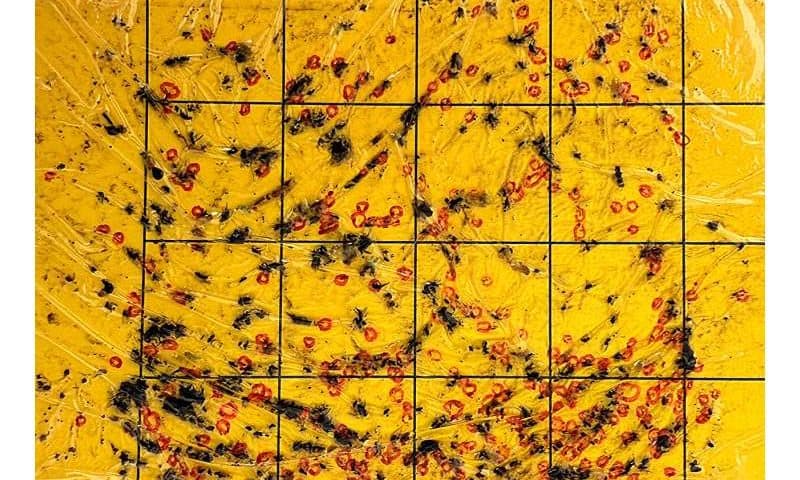
The team analyzed data from nearly 1,700 yellow sticky traps deployed weekly in both open fields and high tunnel production systems for tomatoes and peppers at the Texas A&M AgriLife Research Station at Bushland. Those counts were combined with up to 16 environmental variables, including temperature, humidity, wind speed, wind direction and rainfall, as well as the size of the “parent population” recorded 14 days earlier.
Machine learning models proved highly accurate in predicting pest population development. Models predicted thrips populations in open field settings with nearly 88% accuracy and reached about 85% accuracy in high tunnels.
“AI represents a powerful addition to our modeling because it allows us to analyze many more environmental and biological variables simultaneously and uncover patterns we simply could not see before,” Gadhave said. “This study showed that we can produce very accurate localized forecasts for pest development.”
From reactive to predictive pest management
Gadhave said accuracy dropped sharply in models that applied parameters across both open field and high tunnel systems at the same location. This finding highlights how microclimates function as distinct pest ecosystems, even when fields are side by side.
“What stood out was how quickly models broke down across the different systems,” he said. “Even neighboring fields behaved like different ecosystems, which tells us pest dynamics are fundamentally shaped by microclimate.”
The team also found that one outbreak parameter stood out in both open field and high tunnel systems: parent population size.
If thrips were already present two weeks earlier, the risk of a severe outbreak increased substantially. Temperature ranked next, with wind and humidity shaping how populations spread and build.
This use of AI modeling to forecast localized pest population dynamics shows the potential for its application across various crops, pests and regional microclimates, Gadhave said. Advanced pest and disease forecasting tools could significantly alter how producers monitor and protect their crops.
“This is proof that AI-enabled tools for agriculture aren’t futuristic,” Gadhave said. “They’re already here, and AgriLife Research is well positioned to lead their development and application in the field where they can benefit producers.”