Synbio Launches AI Facial-Analysis Trial Aimed at Earlier Detection of Mental Health Disorders
New proof-of-concept study could pave the way for objective, non-invasive screening for PTSD and depression
Synbio International Inc. (OTC: SYIN) has taken an important step toward transforming how mental health conditions may be identified, announcing a new proof-of-concept clinical trial for an artificial intelligence–based facial analysis technology designed to help screen for disorders such as post-traumatic stress disorder (PTSD) and major depressive disorder (MDD).
The company has signed a Master Services Agreement with CRO Services Pty Ltd, a leading Australian clinical research organization and a subsidiary of ASX-listed Resonance Health Ltd. Under the agreement, CRO Services will conduct a real-world clinical study to evaluate the effectiveness of Synbio partner FacialDx Incorporated’s NIMS™ (Non-invasive Medical Screening) technology.
How the Technology Works — In Simple Terms
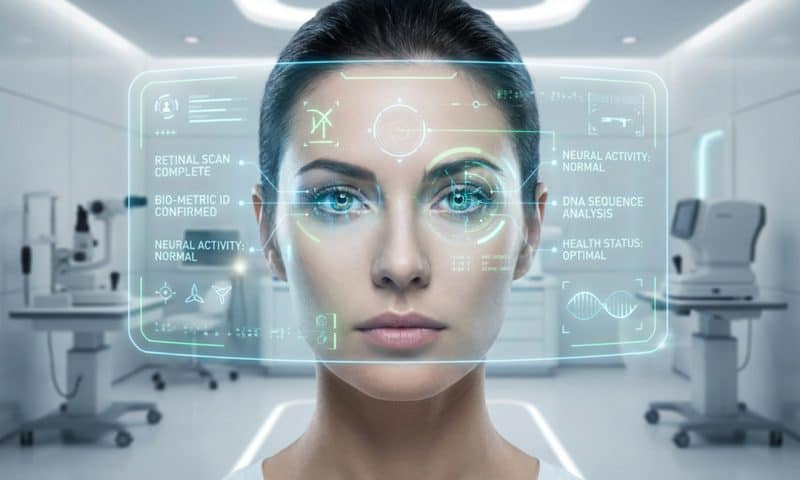
The FacialDX NIMS™ platform uses artificial intelligence to analyze subtle patterns in facial images that may be associated with underlying mental health conditions. Unlike traditional screening methods that rely heavily on patient questionnaires and self-reporting, the technology looks for biological signals expressed in facial features—signals that may not be consciously controlled or easily articulated by patients.
The process is quick, non-invasive, and does not require blood tests, wearable devices, or lengthy interviews. A facial image is captured and analyzed by AI algorithms trained to detect patterns that have been linked, through research, to mental health conditions such as PTSD and depression.
Importantly, the technology is intended to support clinicians, not replace them. The goal is to provide an objective data point that can help clinicians decide when further evaluation or follow-up may be warranted.
Why This Matters
Mental health is already discussed or assessed in an estimated 150 million clinical encounters every year in the United States alone. Yet screening today remains largely subjective, depending on patient honesty, memory, comfort level, and clinician interpretation. Stigma and time constraints can further limit early identification.
If validated through clinical trials, Facial DX’s NIMS™ could become one of the first objective screening tools for mental health conditions—something long viewed as a major unmet need in healthcare. Earlier identification may allow for earlier intervention, potentially improving patient outcomes and reducing long-term societal and economic costs.
Beyond healthcare settings, the technology could also be applied in corporate wellness programs and high-risk industries, where early detection of mental health challenges may improve safety, productivity, and overall well-being.
Strategic Opportunity for Synbio
The proof-of-concept trial represents a critical milestone for Synbio. Clinical validation is a prerequisite for regulatory engagement and eventual commercialization, and conducting the study in Australia allows the company to benefit from efficient timelines and internationally recognized clinical standards.
Success in this trial could position Synbio to pursue commercialization across multiple large markets, including primary care, behavioral health, psychiatry, occupational health, and corporate wellness. The scalability of software-based AI solutions also opens the door to broad deployment without the infrastructure costs associated with traditional diagnostic tools.
Synbio’s leadership views the trial as a foundational step toward building long-term value. With depression now among the leading causes of disability in working-age adults, demand for accessible, objective mental health screening tools continues to grow.
What’s Next
The clinical trial is expected to begin in early 2026 and conclude later in the year. Data generated will be used to guide regulatory strategy and commercial planning. While further agreements and approvals remain pending, the study marks a meaningful move from concept toward real-world clinical validation.
For Synbio, the initiative underscores its broader mission: leveraging clinically validated artificial intelligence to bridge the gap between wellness and medicine—starting with one of healthcare’s most pressing and underserved challenges.
SOURCE: Synbio International